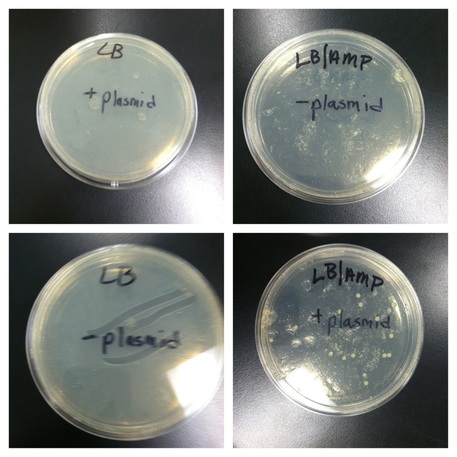
Data and Analysis for pGREEN:
1. Predict your results.
2. Observe the colonies through the petri plate lids. Do not open the plates.
3.
The observed results did not differ from the predictions. The experiment had "text book results."
4. Count the number of individual colonies - if the cell growth is too dense to count individual colonies, record, "lawn."
LB + plasmid (positive control): lawn
LB - plasmid (positive control): lawn
LB/Amp + plasmid (experimental): 23 colonies
LB/Amp - plasmid (negative control): NO growth
5. Compare and contrast the number of colonies on each of the following pairs of plates. What does each pair of results tell you about the experiment?
a. LB + plasmid and LB - plasmid: Both of these plates had lawn growth. This is because the bacteria was free to grow with there being only food. The growth was expected.
b. LB / Amp - plasmid and LB - plasmid: The LB/amp-plasmid had no growth but the LB - plasmid had lawn growth. It was expected that the LB-plasmid would have lawn growth, and the LB/amp - would not because it didn't contain ampicillin resistant gene.
c. LB / Amp + plasmid and LB / Amp - plasmid: The LB / Amp + plasmid did have growth because it was ampicillin-resistant, whereas the LB / Amp - plasmid did not have growth because it was not ampicillin-resistant.
d. LB / Amp + plasmid and LB + plasmid: Both of these showed bacteria growth. The LB/amp + plasmid was ampicillin resistant and grew 23 colonies. The LB + plasmid was also only food, so the bacteria grew.
6. What are you selecting for in this experiment?
Ampicillin resistance is being selected for in this experiment. You test the plasmid for ampicillin resistance by determining whether or not it has been taken up by the bacteria, by subjecting the bacteria to ampicillin in a petri dish. If the bacteria grows, then you know the gene took the ampicillin resistance. If the bacteria died, the bacteria did not take the gene for ampicilin resistance.
7. What does the phenotype of the transformed colonies tell you?
The phenotype of the transformed colonies tells you if bacteria survived or not. If the plate is clean, then there was no bacteria growth. That is because the bacteria was killed by the ampicillin or you did not add bacteria to the dish. If the plate has tiny white dots, then the bacteria survived.
8. What one plate would you first inspect to conclude that the transformation occurred successfully? Why?
The LB/amp + plasmid plate would be the first plate to inspect. This is because it contains the bacteria with the ampicillin resistance plasmid. If the experinment occurred sucessfully the bacteria will grow because it has accepted the plasmid and now is resistant to the ampicilin. However, if the experiment fails, there would not be any bacterial growth. The bacteria would have been killed by the ampicillin.
9. Transformation efficiency is expressed as the number of antibiotic-resistant colonies per ug of plasmid DNA. The object is to determine the mass of plasmid that was spread on the
experimental plate and that was, therefore, responsible for the transformants (number of colonies) observed.
a. Determine the total mass (in μg) of plasmid used. Remember, you used 10 uL of plasmid at a concentration of 0.005 ug / uL.
Total Mass = Volume x Concentration
Total Mass = 10 uL x 0.005 ug / uL
Total Mass = 0.05 μg
b. Calculate the total volume of cell suspension prepared
250µL of calcium chloride + 250µL of luria broth + 10 µL plasmid = 510 µL cell suspension
c. Now calculate the fraction of the total cell suspension that was spread on the plate.
(Volume suspension spread) / (Total volume suspension) = fraction spread
(100 µL) / (510 µL) = 10/51 µL
d. Determine the mass of the plasmid in the cell suspension spread.
[Total mass plasmid (a)] x [fraction spread (c)] = mass plasmid DNA spread
[0.05 μg] x [10/51µL] = 3.921
e. Determine the number of colonies per μg plasmid DNA. Express in scientific notation.
In the LB / AMP + plasmid plate:
Colonies observed / mass plasmid spread (d) = transformation efficiency
1 / 3.921 = 2.55 x 10 ^ -1
10. What factors might influence transformation efficiency?
Antibiotic Concentration: If the concentration is higher, then it may kill more bacteria. If the concentration is lower, it could kill less bacteria.
pH: This can hinder bacteria growth or speed up the growth of the bacteria, depending on whether it is acidic or basic or on the bacteria's optimal pH conditions.
Temperature: The growth of bacteria could be dependent on the temperature. High temperature v. Low temperature.
The observed results did not differ from the predictions. The experiment had "text book results."
4. Count the number of individual colonies - if the cell growth is too dense to count individual colonies, record, "lawn."
LB + plasmid (positive control): lawn
LB - plasmid (positive control): lawn
LB/Amp + plasmid (experimental): 23 colonies
LB/Amp - plasmid (negative control): NO growth
5. Compare and contrast the number of colonies on each of the following pairs of plates. What does each pair of results tell you about the experiment?
a. LB + plasmid and LB - plasmid: Both of these plates had lawn growth. This is because the bacteria was free to grow with there being only food. The growth was expected.
b. LB / Amp - plasmid and LB - plasmid: The LB/amp-plasmid had no growth but the LB - plasmid had lawn growth. It was expected that the LB-plasmid would have lawn growth, and the LB/amp - would not because it didn't contain ampicillin resistant gene.
c. LB / Amp + plasmid and LB / Amp - plasmid: The LB / Amp + plasmid did have growth because it was ampicillin-resistant, whereas the LB / Amp - plasmid did not have growth because it was not ampicillin-resistant.
d. LB / Amp + plasmid and LB + plasmid: Both of these showed bacteria growth. The LB/amp + plasmid was ampicillin resistant and grew 23 colonies. The LB + plasmid was also only food, so the bacteria grew.
6. What are you selecting for in this experiment?
Ampicillin resistance is being selected for in this experiment. You test the plasmid for ampicillin resistance by determining whether or not it has been taken up by the bacteria, by subjecting the bacteria to ampicillin in a petri dish. If the bacteria grows, then you know the gene took the ampicillin resistance. If the bacteria died, the bacteria did not take the gene for ampicilin resistance.
7. What does the phenotype of the transformed colonies tell you?
The phenotype of the transformed colonies tells you if bacteria survived or not. If the plate is clean, then there was no bacteria growth. That is because the bacteria was killed by the ampicillin or you did not add bacteria to the dish. If the plate has tiny white dots, then the bacteria survived.
8. What one plate would you first inspect to conclude that the transformation occurred successfully? Why?
The LB/amp + plasmid plate would be the first plate to inspect. This is because it contains the bacteria with the ampicillin resistance plasmid. If the experinment occurred sucessfully the bacteria will grow because it has accepted the plasmid and now is resistant to the ampicilin. However, if the experiment fails, there would not be any bacterial growth. The bacteria would have been killed by the ampicillin.
9. Transformation efficiency is expressed as the number of antibiotic-resistant colonies per ug of plasmid DNA. The object is to determine the mass of plasmid that was spread on the
experimental plate and that was, therefore, responsible for the transformants (number of colonies) observed.
a. Determine the total mass (in μg) of plasmid used. Remember, you used 10 uL of plasmid at a concentration of 0.005 ug / uL.
Total Mass = Volume x Concentration
Total Mass = 10 uL x 0.005 ug / uL
Total Mass = 0.05 μg
b. Calculate the total volume of cell suspension prepared
250µL of calcium chloride + 250µL of luria broth + 10 µL plasmid = 510 µL cell suspension
c. Now calculate the fraction of the total cell suspension that was spread on the plate.
(Volume suspension spread) / (Total volume suspension) = fraction spread
(100 µL) / (510 µL) = 10/51 µL
d. Determine the mass of the plasmid in the cell suspension spread.
[Total mass plasmid (a)] x [fraction spread (c)] = mass plasmid DNA spread
[0.05 μg] x [10/51µL] = 3.921
e. Determine the number of colonies per μg plasmid DNA. Express in scientific notation.
In the LB / AMP + plasmid plate:
Colonies observed / mass plasmid spread (d) = transformation efficiency
1 / 3.921 = 2.55 x 10 ^ -1
10. What factors might influence transformation efficiency?
Antibiotic Concentration: If the concentration is higher, then it may kill more bacteria. If the concentration is lower, it could kill less bacteria.
pH: This can hinder bacteria growth or speed up the growth of the bacteria, depending on whether it is acidic or basic or on the bacteria's optimal pH conditions.
Temperature: The growth of bacteria could be dependent on the temperature. High temperature v. Low temperature.